A Quest for Reproducible Data Science
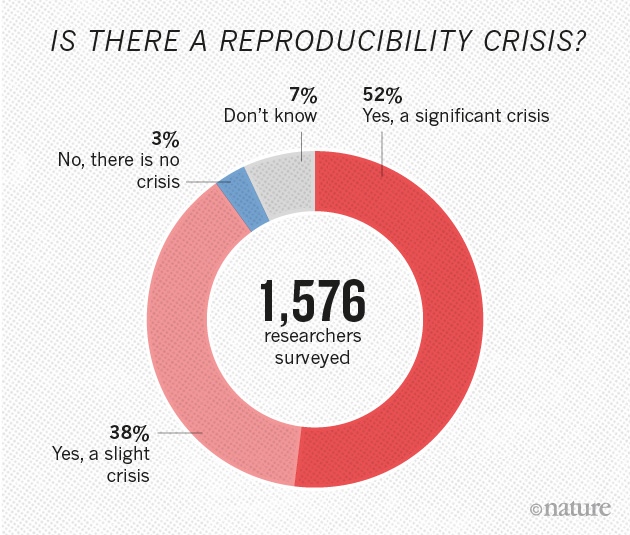 Figure from Nature’s survey of 1,576 researchers who took a brief online questionnaire on reproducibility in research.
Figure from Nature’s survey of 1,576 researchers who took a brief online questionnaire on reproducibility in research.Have you ever tried to reproduce someone else’s analysis and got different results? Ever tried to repeat a lab experiment and failed? If yes, you are not alone. More than 70% of researchers tried and failed to reproduce another scientist’s experiment.
In Nature’s online survey from 2016, more than half of participants agreed that there is a significant reproducibility crisis and 38% percent answered that there is a slight reproducibility crisis. We could argue that people who clicked to the questionnaire about reproducibility are more aware of those concerns, but don’t we all agree that science should be open and reproducible?
According to metascience – the scientific study of science itself – replication is an essential part of the scientific method. It increases the quality of scientific research and reduces waste. Another important aspect of science is scientific creativity, the ability to see novel and unexpected patterns in data, but sometimes random noise can look just like a pattern that fits our desires. This poses a danger to the true discovery rate leading to the accumulation of incorrect findings and wasting future research efforts.
In A manifesto for reproducible science , authors argue for the adoption of measures to optimise key elements of the scientific process: methods, reporting and dissemination, reproducibility, evaluation and incentives. The goals of their measures are improving the transparency, reproducibility, and efficiency of scientific research. Examples of potential solutions are: rigorous training in statistics and research methods for future researchers, use of reporting and protocol checklists, open data, materials and software, pre- and post- publication peer review, disclosure of conflicts of interest, and funding replication studies.

After reading this manifesto, I asked myself: what can I do to fight the reproducibility crisis? I definitely can strive to write reproducible code while documenting all the software, tools, and parameters used. The use of general-purpose workflow languages ( CWL , Nextflow , and Snakemake ) or domain-specific ones ( NGLess , language for next-generation sequencing data processing) facilitates the creation of reproducible, scalable, and portable data analyses. Such tools are designed to meet the needs of data-intensive sciences by providing the framework for reproducible analysis. Workflows can be easily scaled to server, cluster or cloud, and software required to run the analysis are automatically deployed to any execution environment.
Imagine someone told you that you could throw away all the intermediate files because as long as you have raw data, you could get these files anytime. To quote Luis Pedro Coelho, the main developer of NGLess:
Results are equivalent to data, just given more computation.
So let’s make even our supplementary material reproducible!